Research Interests
Lab Interest in Photoreceptor Degeneration
The research focus of my group is on photoreceptor metabolism and the signaling pathways that regulate photoreceptor metabolism. We study retinal degenerative diseases that affect cone photoreceptors, since cones are essential for color, daylight and high acuity vision in humans. However, because of the peculiar interdependence between rods and cones in humans and mouse we are also interested in rod photoreceptors. In particular, we are interested in how cell metabolism is controlled in both types of photoreceptors of healthy retinas and how cell metabolism adapts during diseases that cause photoreceptor loss. Another area of interest is how photoreceptor metabolism adapts during the process of aging, since aging is known to lead to a host of metabolic changes in the entire body. Additionally, metabolic disease conditions that affect the entire body such as diabetes and only secondarily cause retinal abnormalities such as diabetic retinopathy (DR) are also of interests.
The reason why we are interested in understanding how photoreceptor metabolism is regulated is that photoreceptors are among the highest energy consuming cells in the human body1. Two circumstances contribute to the fact that photoreceptors have such a high-energy demand. First, like all neurons photoreceptors need large quantities of ATP in order to re-equilibrate membrane potential1. Second, photoreceptors are constantly growing cells yet they do not divide. Photoreceptors need to synthesize every day membranes and proteins they lose due to the shedding of their outer segments2. The photoreceptor outer segment is so densely packed2-4 to optimize absorption of light photons, that the average lipid content of a photoreceptor is 15% of its cell mass compared to 1% for normal cells5,6 and each photoreceptor contains roughly 60pg of protein6,7. Since photoreceptors shed 10% of their OS daily the lipid and protein content that needs to be re-synthesized amounts roughly to that of a cell division per day, suggesting that photoreceptors should have a metabolic profile similar to that of proliferating cells.
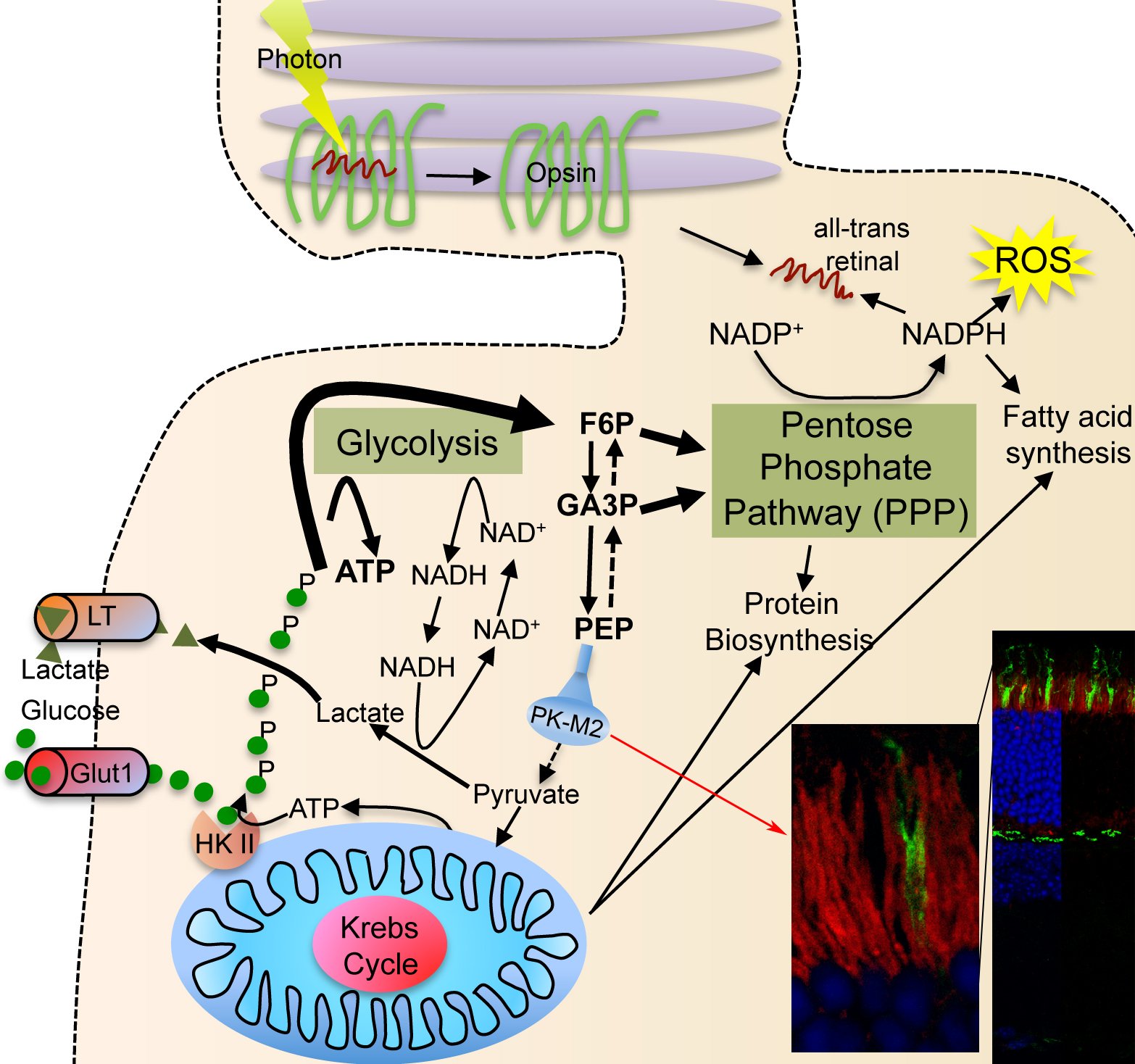 |
| Photoreceptor Metabolism: Glucose is diverged into the PPP by expression of PKM2 (red staining in section showing rod inner segments). The PPP allows for more NADPH synthesis, which is important for reactive oxygen species neutralization, fatty acid synthesis for outer segment growth and chromophore recycling. |
Proliferating cells use a lot of glucose to meet their anabolic needs8. Cell anabolism is accomplished by diverging large quantities of glycolytic intermediates into the oxidative and non-oxidative branch of the pentose phosphate pathway (PPP). The divergence of glycolytic intermediates into the PPP is achieved by expression of the embryonic M2 splice isoform of pyruvate kinase (PKM2), which reduced activity acts as a bottleneck allowing accumulated intermediate products of glycolysis to be shunt into PPP9,10. This is accompanied by production and secretion lactate in order to regenerate NAD+ that is needed to maintain the high metabolic flux. Secretion of lactate in the presence of ample oxygen is a phenomenon commonly referred to as Warburg effect8. To efficiently trap large quantities of glucose proliferating cells express hexokinase II, which is absent from most tissues11. This isoform of hexokinase phosphorylates and traps intracellular glucose more efficiently12. All these metabolic adaptations, which allow cells to proliferate, are regulated in large by the kinase mechanistic target of rapamycin (mTOR), which controls the expression of a gene regulatory network including PKM2, glycolysis, the PPP, and lipid biosynthesis13.
Similar to proliferating cells, most of the glucose taken up by photoreceptors does not enter the Krebs cycle14. Photoreceptors secrete large amounts of lactate14-16 although they are exposed to almost arterial concentrations of oxygen17. Additionally, photoreceptors express also hexokinase II and PKM2 in their inner segments18, where the glucose transporters are located19. The similarity between photoreceptors metabolism and that of proliferating cells thus attributes a central role to mTOR in photoreceptors. Therefore, the goal of the lab is thus to understand how the insulin/mTOR pathway regulates cell metabolism in photoreceptors and how it adapts photoreceptor metabolism to various retinal and systemic metabolic diseases.
Bibliography
1. Ames, A., 3rd. CNS energy metabolism as related to function. Brain Res Brain Res Rev 34, 42-68 (2000).
2. Young, R.W. The renewal of rod and cone outer segments in the rhesus monkey. J Cell Biol 49, 303-318 (1971).
3. Bownds, D., Gordon-Walker, A., Gaide-Huguenin, A.C. & Robinson, W. Characterization and analysis of frog photoreceptor membranes. J Gen Physiol 58, 225-237 (1971).
4. Lisman, J.E. & Bering, H. Electrophysiological measurement of the number of rhodopsin molecules in single Limulus photoreceptors. J Gen Physiol 70, 621-633 (1977).
5. Whikehart, D.R. Biochemistry of the Eye, 2nd Edition, (2003).
6. Scott, B.L., Racz, E., Lolley, R.N. & Bazan, N.G. Developing rod photoreceptors from normal and mutant Rd mouse retinas: altered fatty acid composition early in development of the mutant. J Neurosci Res 20, 202-211 (1988).
7. Lowry, O.H., Rosebrough, N.J., Farr, A.L. & Randall, R.J. Protein measurement with the Folin phenol reagent. J Biol Chem 193, 265-275 (1951).
8. Vander Heiden, M.G., Cantley, L.C. & Thompson, C.B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029-1033 (2009).
9. Vander Heiden, M.G., et al. Evidence for an alternative glycolytic pathway in rapidly proliferating cells. Science 329, 1492-1499 (2010).
10. Christofk, H.R., et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230-233 (2008).
11. Cardenas, M.L., Cornish-Bowden, A. & Ureta, T. Evolution and regulatory role of the hexokinases. Biochim Biophys Acta 1401, 242-264 (1998).
12. Mathupala, S.P., Ko, Y.H. & Pedersen, P.L. Hexokinase II: cancer's double-edged sword acting as both facilitator and gatekeeper of malignancy when bound to mitochondria. Oncogene 25, 4777-4786 (2006).
13. Duvel, K., et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol Cell 39, 171-183 (2010).
14. Winkler, B.S., Starnes, C.A., Sauer, M.W., Firouzgan, Z. & Chen, S.C. Cultured retinal neuronal cells and Muller cells both show net production of lactate. Neurochem Int 45, 311-320 (2004).
15. Winkler, B.S., Pourcho, R.G., Starnes, C., Slocum, J. & Slocum, N. Metabolic mapping in mammalian retina: a biochemical and 3H-2-deoxyglucose autoradiographic study. Exp Eye Res 77, 327-337 (2003).
16. Chertov, A.O., et al. Roles of glucose in photoreceptor survival. J Biol Chem 286, 34700-34711 (2011).
17. Bill, A., Sperber, G. & Ujiie, K. Physiology of the choroidal vascular bed. Int Ophthalmol 6, 101-107 (1983).
18. Reidel, B., et al. Proteomic profiling of a layered tissue reveals unique glycolytic specializations of photoreceptor cells. Mol Cell Proteomics 10, M110 002469 (2011).
19. Gospe, S.M., 3rd, Baker, S.A. & Arshavsky, V.Y. Facilitative glucose transporter Glut1 is actively excluded from rod outer segments. J Cell Sci 123, 3639-3644 (2010).
