-

-

BUCCI
-
COLUBRI
-

SASSETTI
-

JACOBSON
-

ZHUGE
-

IRAZOQUI
Research Labs in Microbiology
 |
Behar We seek to understand how the causative agent of tuberculosis, namely the bacterium Mycobacterium tuberculosis evades host immunity, and how different components of the immune response work together to contain infection. Our ultimate goal is to inform vaccine development and the design of host directed therapy. |
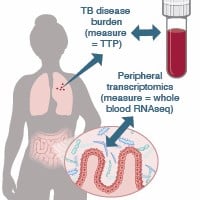 |
Bucci The Bucci Lab seek to understand mechanisms responsible for the temporal and spatial dynamics of complex biological systems such as the human microbiome and to develop tools to predict and engineer them. |
 |
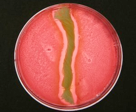
Colubri The Colubri Lab (CoLabo) conducts research at the interface between genetics, epidemiology, machine learning, and data visualization. The goal is to create new tools that integrate heterogenous data sources (ranging from genome sequences to environmental factors) with applications not only in infectious disease prediction but also in science outreach and STEM education. |
 |
Durham |
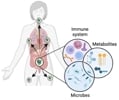 |
Galván Peña We investigate how intestinal immune populations integrate signals from the microbiota and communicate with other cell populations and microbial metabolites at distal tissues. |
 |
Gerstein |
 |
He Control of eukaryotic mRNA translation and decay. |
 |
Irazoqui Host-Microbe Interactions, including pathogenic bacteria, viruses, and microbiome. Brain-gut-microbiota axis in health and disease. SARS-CoV-2 cellular pathogenesis. |
 |
Jacobson Cytoplasmic aspects of post-transcriptional regulation in eukaryotes. |
 |
Kowalik Cellular and organismal studies of viral infections. |
 |
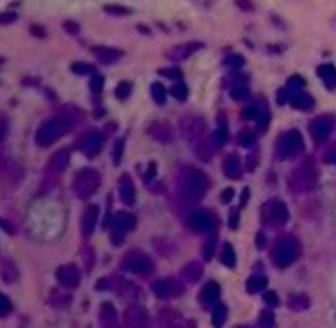
Maldonado-Contreras We investigate how manipulation of the gut microbiota through diet can improve clinical outcomes in patients with chronic diseases. |
 |
McCormick We investigate epithelial barrier function in the intestine and mucosal inflammation during enteric bacterial pathogen infection and inflammatory bowel diseases. |
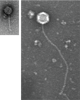 |
McKitterick The McKitterick lab focuses on (1) how the mycomembrane and other layers of the envelope are assembled and disassembled using phages to guide new discoveries and (2) isolating and characterizing new phages that infect mycolated bacteria. |
 |
Morrison We study basic mechanisms of infection of enveloped, RNA viruses as well as vaccine development. |
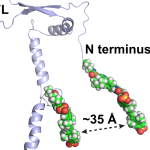 |
Munro We are devoted to the development and application of quantitative, biophysical approaches to studying a variety viral pathogens, including HIV, influenza, Ebola, Lassa, and SARS-CoV-2. |
 |
Murphy My lab focuses on recombineering technologies for the modification of mycobacterial genes, identification of new antibiotics for tuberculosis, and the molecular mechanisms of mycobacterial DNA mismatch repair. |
 |
Papavinasasundaram |
 |
Sassetti The development of transformational new interventions for tuberculosis relies on understanding the fundamental biology that determines the outcome of an infection. The Sassetti lab applies a combination of mammalian and bacterial genetic strategies to understand the intricate interactions between phenotypically diverse host and pathogen influence pathogenesis and the response to therapy. |
 |
Schrader B cell immunology focusing on antibody diversification mechanisms of class switch recombination and somatic hypermutation. |
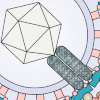 |
Tai The Tai lab works towares the development the next-generation of rAAVs and other therapeutic platforms. |
 |
Ward Our group applies microbial genomic and metagenomic approaches to understand the how bacteria contribute to human heath and infectious disease, focusing on the transmission of pathogenic strains both in the hospital and in the community. |
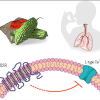 |
Zhuge The ZhuGe lab studies the molecular mechanisms by which Ca2+ signals and ion channels control neurotransmitter release and smooth muscle contractility. We also exploit new physiological functions of bitter taste receptors in extra-oral tissues and organs. |
-

-

BUCCI
-

COLUBRI
-

SASSETTI
-

JACOBSON
-

ZHUGE
-

IRAZOQUI