The UPRmt and the propagation of deleterious mitochondrial genomes.
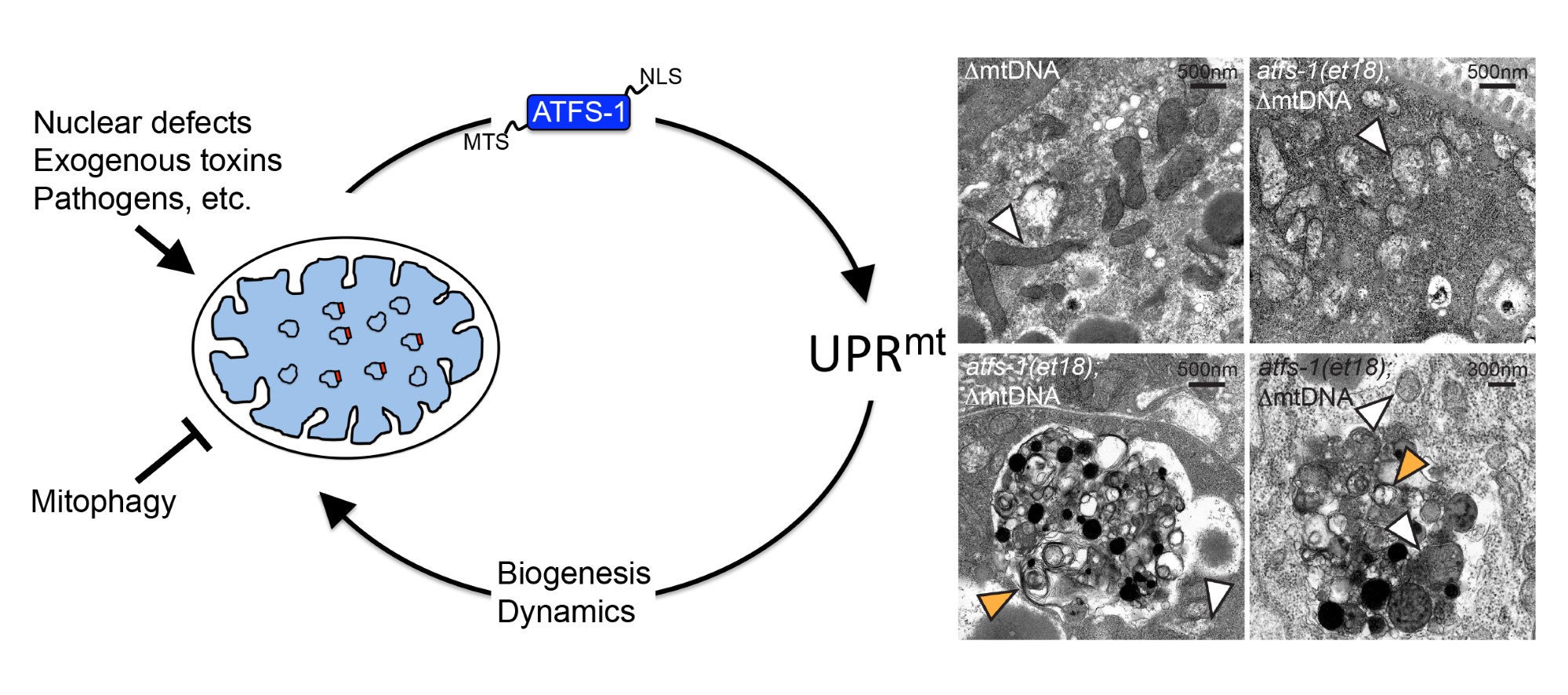
Most cells harbor hundreds of mitochondrial genomes (mtDNA) that encode thirteen essential components of the respiratory chain and ATP synthase. And, mutations or deletions in a low percentage of mtDNAs are well-tolerated. However, if the percentage of deleterious mtDNAs reaches ~70%, the ensuing mitochondrial dysfunction can result in disease or pathology. One means to reach this percentage is through maternal inheritance. Alternatively, as we age differentiated, post-mitotic cells such as muscles and neurons accumulate deleterious mtDNAs that perturb mitochondrial dysfunction though the mechanisms are poorly understood. We recently demonstrated in a C. elegans model that the presence and propagation of a clonal deleterious mtDNA requires ATFS-1 and the UPRmt. 60% heteroplasmy caused constitutive activation of the UPRmt, which was absolutely essential for maintenance and propagation of the deleterious mtDNA1. Interestingly, UPRmt activation was also sufficient to propagate the deleterious mtDNAs to extreme toxicity. Thus, we propose that in the context of mtDNA heteroplasmy, constitutive UPRmt activation3 caused by OxPhos defects propagates or maintains the deleterious mtDNA in an attempt to recover OxPhos activity by promoting mitochondrial biogenesis and dynamics.
1. How do deleterious mtDNAs outcompete wildtype mtDNAs?
2. Do the deleterious mtDNAs that arise during aging or in cancers require the UPRmt?
3. How is the clearance of deleterious mtDNA in the female germline regulated?
4. How is mtDNA transcription and replication regulated during mitochondrial stress2?
References:
1Lin YF, Schulz AM, Pellegrino MW, Lu Y, Shaham S, Haynes CM. (2016) Maintenance and propagation of a deleterious mitochondrial genome by the mitochondrial unfolded protein response. Nature. May 2:533(7603): 416-419. PMID: 27135930
2Nargund AN, Fiorese CJ, Pellegrino MW, Deng P, Haynes CM. (2015) Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPRmt. Molecular Cell. April 2;58(1):123-33. PMID: 25773600
3Lamech, LT, Haynes CM. (2015) The unpredictability of prolonged activation of stress response pathways. Journal of Cell Biology. June 22:209(6): 781-87. PMID: 26101215
