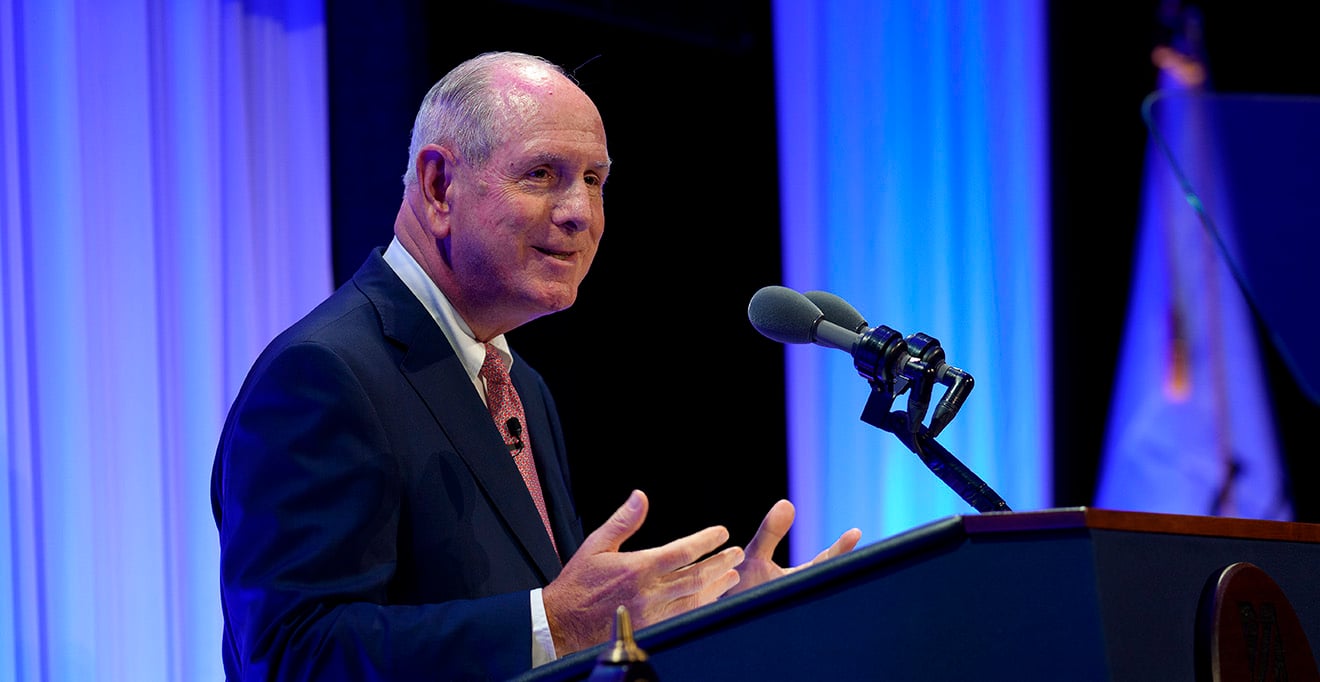
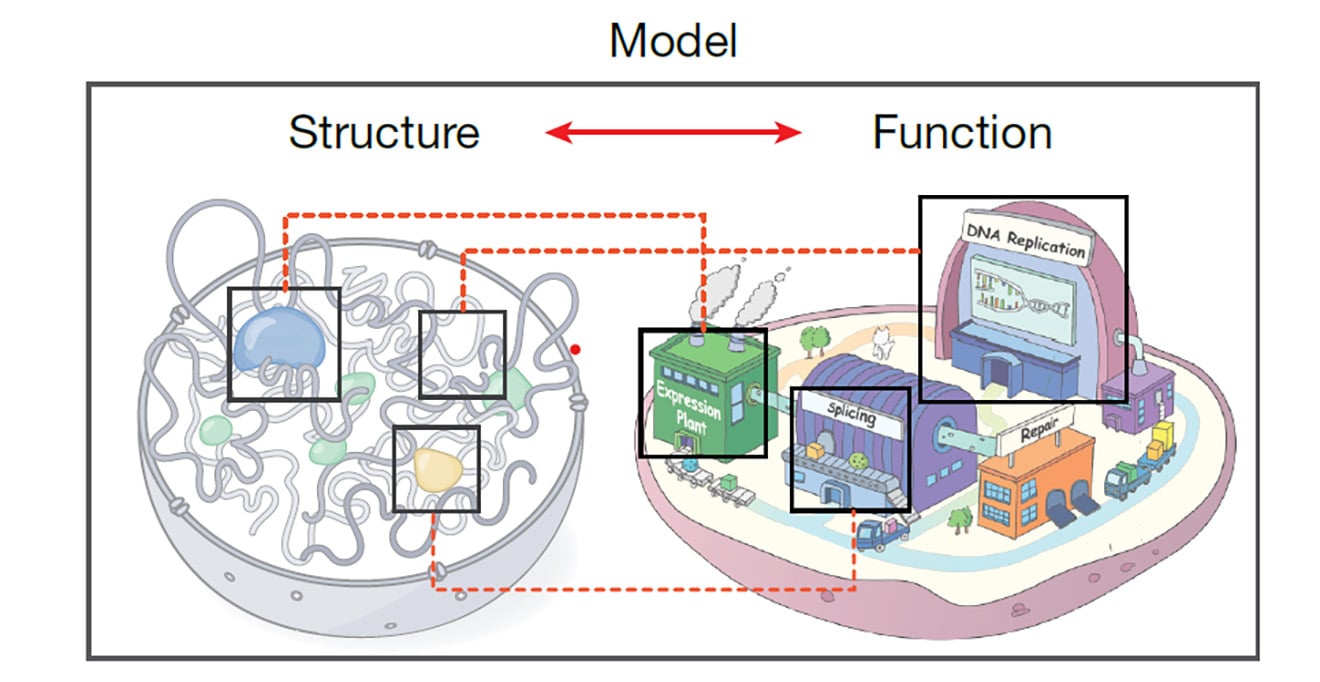
Our studies will contribute to our general understanding of signaling regulatory networks and protein-protein interactions, with relevance to the mechanisms underlying normal cell function as well as disease states.
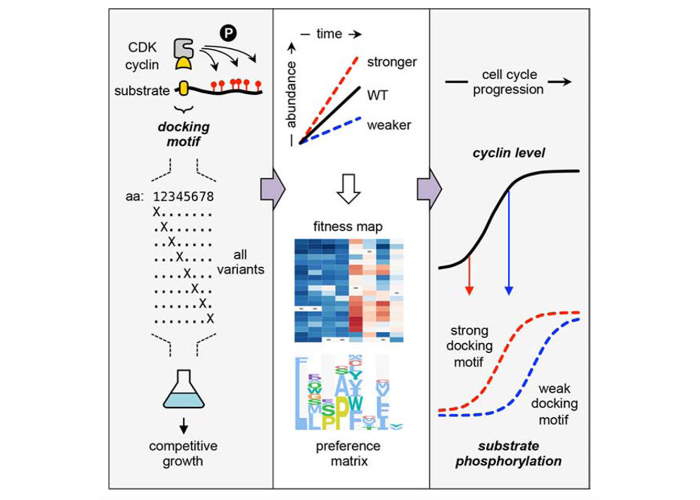 Comprehensive Analysis of G1 Cyclin Docking Motif Sequences that Control CDK Regulatory Potency In Vivo.
Comprehensive Analysis of G1 Cyclin Docking Motif Sequences that Control CDK Regulatory Potency In Vivo.
Bandyopadhyay S, Bhaduri S, Örd M, Davey NE, Loog M, Pryciak PM.
Curr Biol. 2020 Nov 16;30(22):4454-4466.e5. doi: 10.1016/j.cub.2020.08.099. Epub 2020 Sep 24. PMID: 32976810; PMCID: PMC8009629.
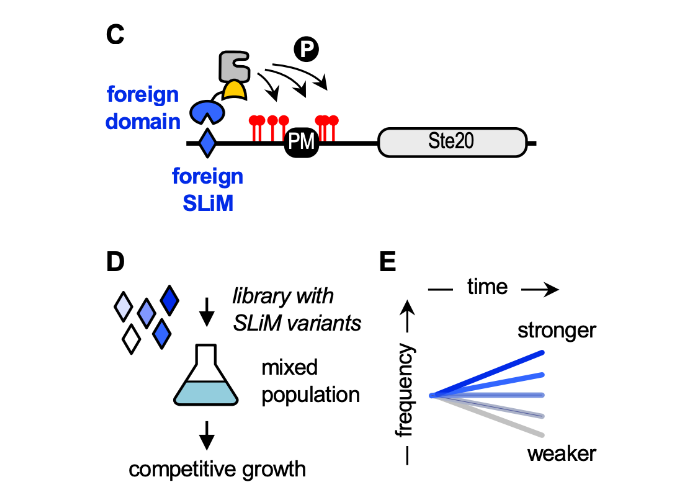 A quantitative intracellular peptide binding assay reveals recognition determinants and context dependence of short linear motifs
A quantitative intracellular peptide binding assay reveals recognition determinants and context dependence of short linear motifs
Mythili S. Subbanna, Matthew J. Winters, Mihkel Örd, Norman E. Davey, and Peter M. Pryciak
bioRxiv 2024.10.30.621084; doi: https://doi.org/10.1101/2024.10.30.621084
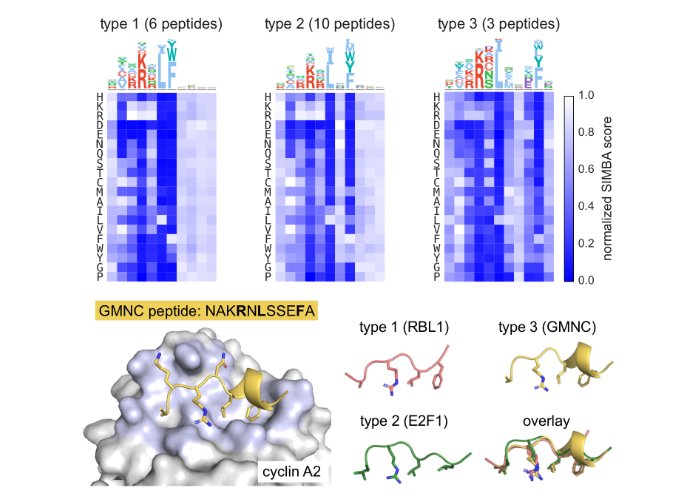 High-throughput discovery and deep characterization of cyclin-CDK docking motifs
High-throughput discovery and deep characterization of cyclin-CDK docking motifs
Mihkel Örd, Matthew J. Winters, Mythili S. Subbanna, Natalia de Martin Garrido, Victoria I. Cushing, Johanna Kliche, Caroline Benz, Ylva Ivarsson, Basil J. Greber, Peter M. Pryciak*, Norman E. Davey*
bioRxiv 2024.12.03.625240; doi: https://doi.org/10.1101/2024.12.03.625240