How Chromosome Structure Influences Nuclear Processes
Over the years, our research has focused on understanding how chromatin dynamics regulates a number of nuclear processes, including transcription, DNA double-strand break repair, and DNA replication. Currently, our efforts are focused exclusively on the role of chromatin in transcriptional regulation. In general, research is focused on two interlocking groups of projects–(1) Biochemical and biophysical analyses of ATP-dependent chromatin remodeling “machines” and (2) Genome-wide approaches in budding yeast that focuses on how chromatin impacts “transcriptional homeostasis”, which in general terms refers to the global regulation of coding and non-coding RNAs as a function of the cell cycle, environmental changes, or mRNA metabolism
How do chromatin remodeling enzymes work?
Chromatin remodeling enzymes are DNA translocases
DNA unwrapping model for histone dimer exchange
Our lab identified and purified one of the first ATP-dependent chromatin remodeling enzymes, the 1.14 MDa SWI/SNF complex (Cote et al., Science, 1995). Over the years, we have shown that this enzyme is recruited to target genes in vivo and in vitro by sequence-specific transcriptional activators (Yudkovsky et al., Genes Dev.., 1999) and that it plays key roles in mitotic gene expression (Krebs et al., Cell, 2000), as well as DNA double-strand break repair (Bennett and Peterson, Nature Communications, 2015). We have also collaborated with several groups to evaluate the impact of a host of purified remodeling enzymes on in vitro DNA replication (Azmi et al, Molecular Cell, 2017) and promoter nucleosome positioning (Korber et al., Cell, 2017).
Our current biochemical studies are focused on the 14 subunit SWR1C complex that uses the energy of ATP hydrolysis to replace nucleosomal histone H2A with the histone variant H2A.Z. We have developed state-of-the-art ensemble and single-molecule FRET-based assays that follow both the eviction of nucleosomal H2A and the deposition of H2A.Z. Our current work indicates that SWR1C uses multiple subunits to anchor itself to nucleosomal histones, whereby it then uses ATP hydrolysis to drive a limited amount of DNA translocation that leads to H2A de-stabilization and eviction.
Maintenance of Transcriptional Homeostasis
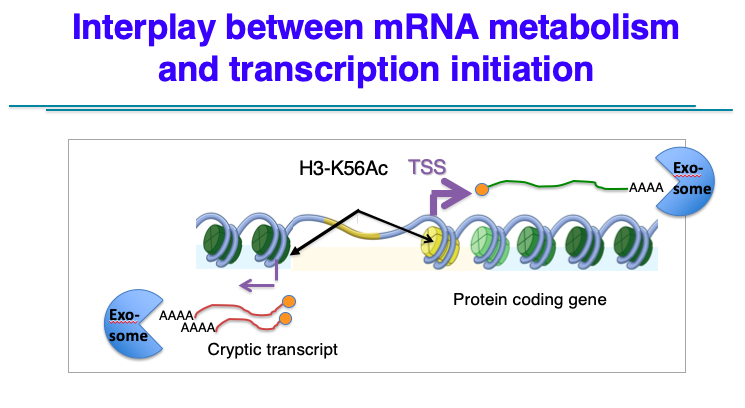
Our in vivo studies are currently focused on two areas that impinge on the concept of general transcriptional homeostasis. First, we are investigating a phenomenon whereby global stabilization of mRNAs (due to inactivation of the mRNA degradation machinery) leads to down-regulation of transcription initiation. For instance, rapid removal of the nuclear RNA exosome both stabilizes many RNAs (mRNAs and ncRNAs) but this also leads to a global decrease in transcription initiation, as assayed by Nascent Transcript Sequencing (Net-seq). Currently, we are investigating a model whereby mRNA stabilization leads to over-expression of selected chromatin regulators that impinge on nucleosome dynamics at promoter regions.
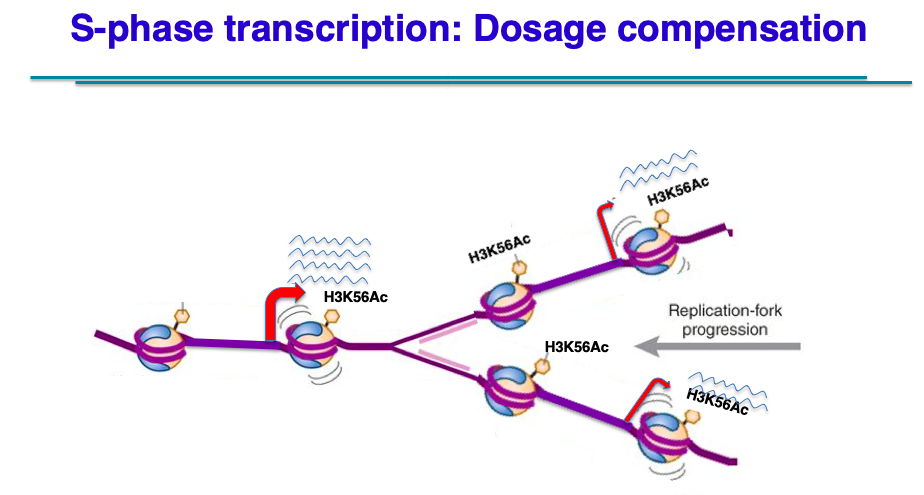
In the second in vivo project, we are investigating a conserved process whereby eukaryotic cells regulate the expression of newly replicated genes so as to control gene expression with cell size. In both yeast and mammalian cells, the duplication of genes during S phase does not lead immediately to a 2x increase in transcription. Rather, gene copies on each sister chromatid are only expressed at 50% levels, leading to an overall G1-level of transcription. This buffering process apparently functions to ensure that gene products (e.g. proteins) are expressed at levels appropriate for cell size. Indeed, as cells enter G2 and cells double their volume, gene transcription shows a global 2x increase in transcription in a step-wise fashion. Essentially, this is a form of gene dosage compensation that is cell cycle regulated. What regulates S phase dosage compensation is unclear, but we are currently entertaining models involving the replication-dependent assembly of cohesion or condensin complexes that might repress transcription in a dosage dependent fashion.
