Our most recent publications:
Total: displaying 10 out of 15 results
In the News
Latest BMB news as reported by the UMass Chan Communications Office
-
-
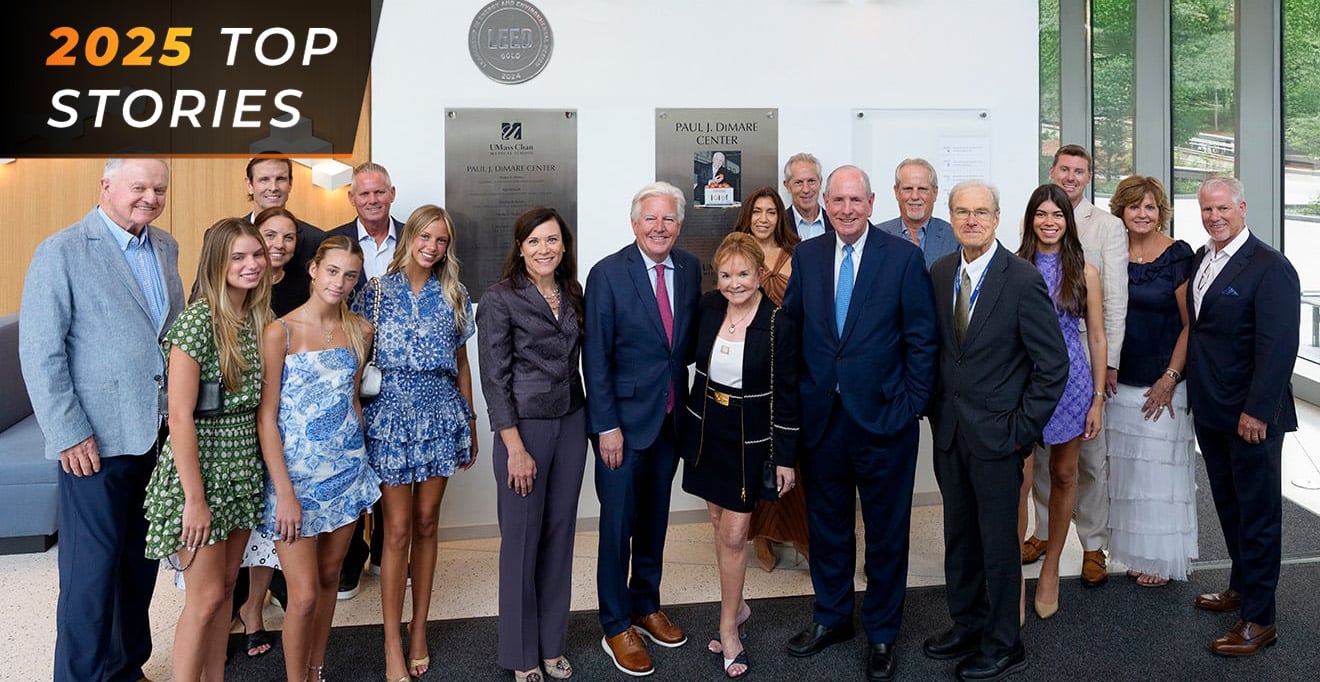
Top story: UMass Chan celebrates naming of Paul J. DiMare Center
December 15, 2025 -

-

-

-
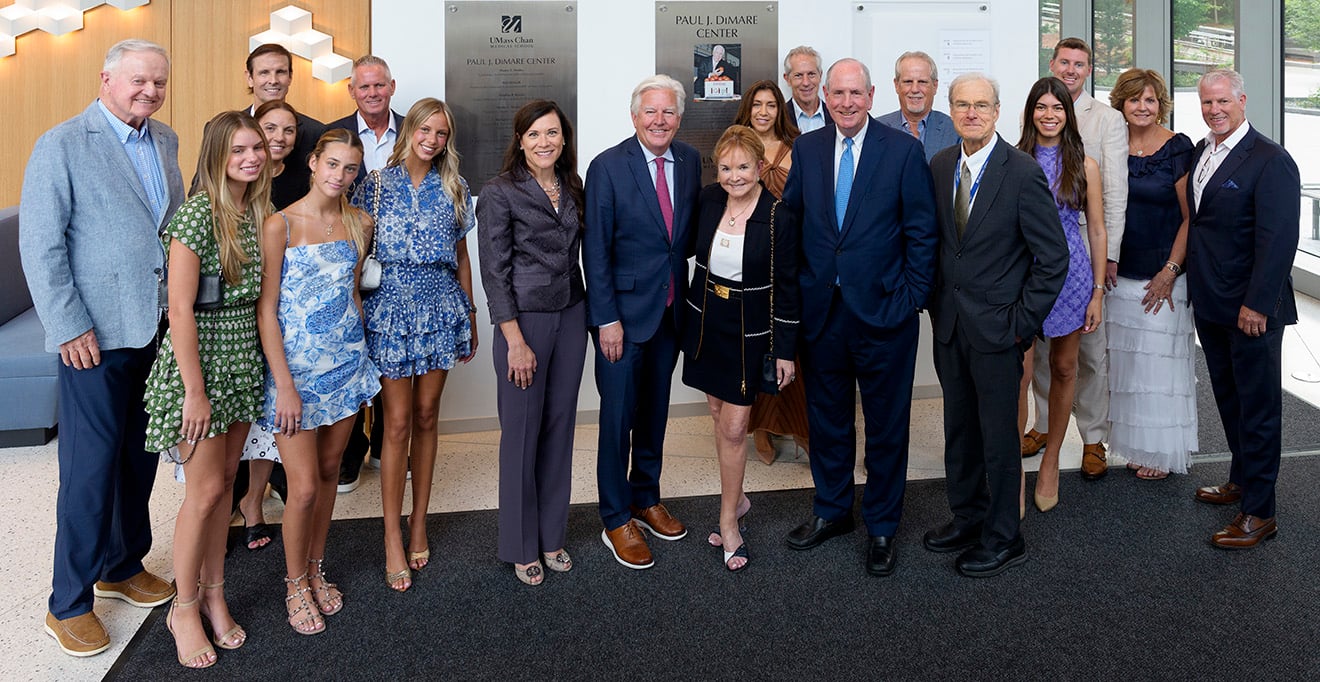

UMass Chan celebrates naming of Paul J. DiMare Center
July 10, 2025 -
-

-

-

-

-

-

-

-

-

Convocation 2024: UMass Chan racing toward ambitious future
September 13, 2024
