Photo: Greg Kahn, Howard Hughes Medical Institute
STAT News, the award winning, Boston-based science and health care digital media outlet, has selected a study on the 4D nucleome led by Job Dekker, PhD, to compete in its annual STAT Madness competition, a bracket-style competition in which readers vote on the most important and impactful research from the previous year. The first round of voting, which will narrow the 2025 field from 64 to 32, begins March 2.
Dr. Dekker, Howard Hughes Medical Institute Investigator, the Joseph J. Byrne Chair in Biomedical Research and professor of systems biology, and an international team of researchers say the work will help scientists decide how to approach specific research questions because the genome’s shape can illuminate what regulatory elements are expressed and how genes work together.
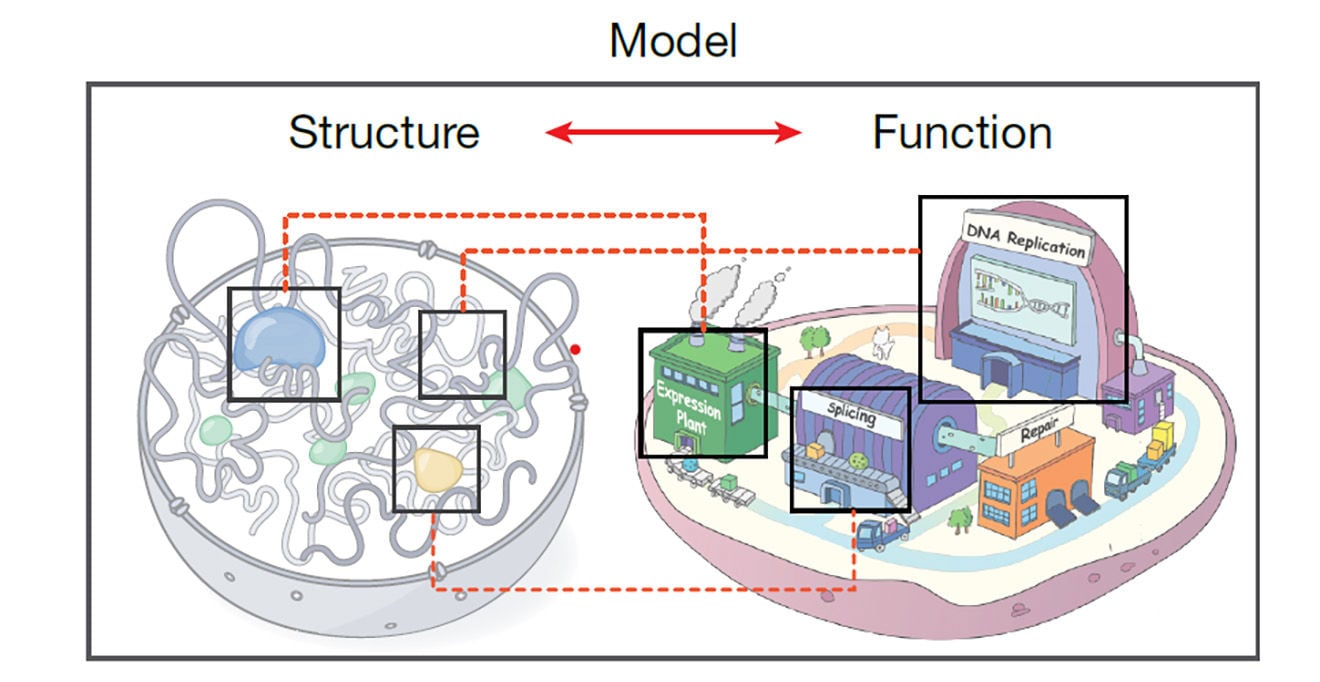
Dekker and colleagues from the 4D Nucleome Consortium (4DN), which he leads, analyzed the genome’s structure to develop an intimate understanding of its folds and loops. Using a dozen experimental methods, along with computational and modeling tools, they created a “scientific user’s guide” that catalogs the role of various assays in determining specific structural elements. Their table and decision tree lay out how each method contributes to a broader understanding of the nucleome’s structure.
“This is the most detailed view of the living physical genome as it exists inside of cells,” said Dekker. “It’s the foundation for the deep exploration of structure and function of the genome.”
The 4DN consortium has, among other findings presented in the study, compiled an extensive catalogue of 140,000 looping interactions between specific elements that no single methodology could have detected on its own. The study also used integrated models to assign dynamic genomic functions comprising multiple genes and regulatory elements, such as replication and transcription, into a three-dimensional context.
The study was published Dec. 15 in Nature.
Members of the public are encouraged to vote for their favorite research. The winning research team will be named in early April after six rounds of voting. To see our coverage of the competition, follow us on X (formerly Twitter), Facebook and LinkedIn.