DNA Polymerase Clamp Loaders
Chromosomal DNA replication requires that DNA polymerases be tethered to ring-shaped sliding clamps that encircle the DNA and allow for high-speed, processive replication. Sliding clamps are loaded onto DNA by the clamp loader complex, a pentameric assembly of proteins of the AAA+ family of ATPases.
We want to understand the function and mechanism of clamp loaders in atomic detail. Recent structures of the T4 bacteriophage clamp loader in complex with primer-template DNA and the sliding clamp have suggested a working model for the clamp loader mechanism as shown below (see Kelch et al Science 2011):
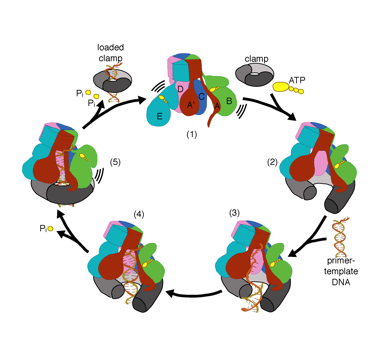
Our model predicts that ATP hydrolysis begins at the end of the AAA+ spiral and proceeds in a sequential fashion up the spiral, resulting in the ejection of the clamp loader from the DNA and the closed clamp. Our hypothesis is illustrated in this animation of three consecutive ATP hydrolysis events:
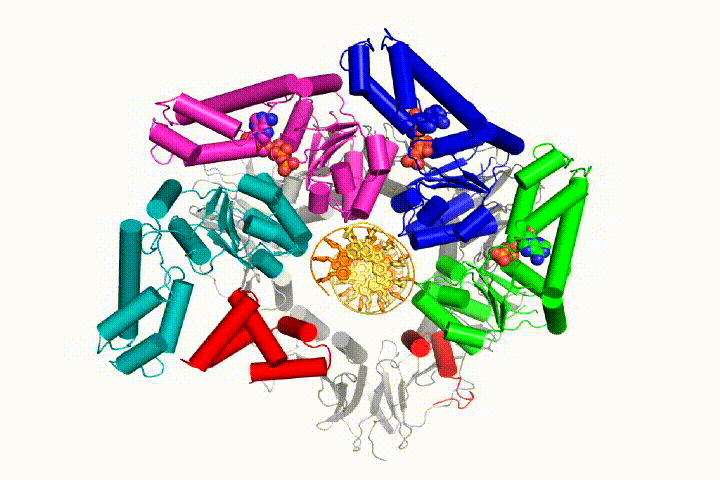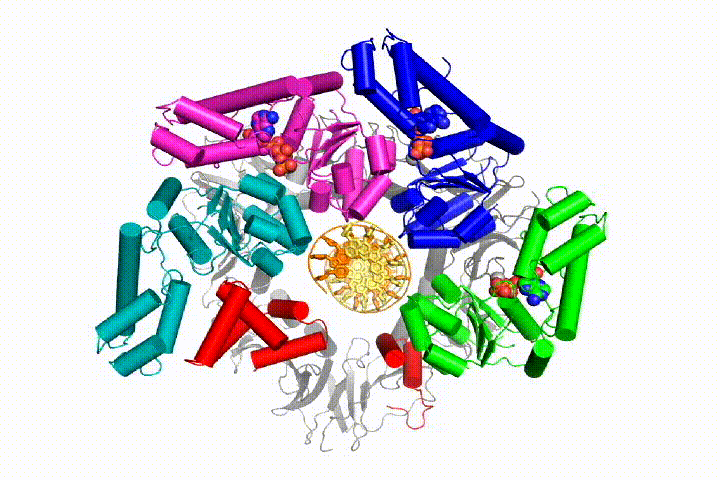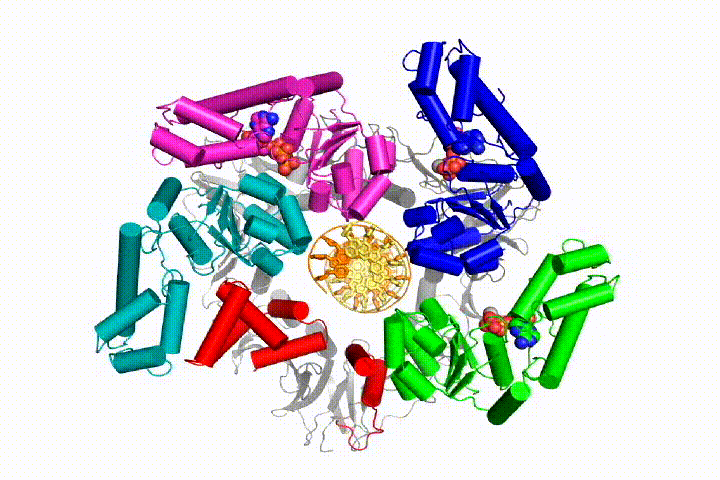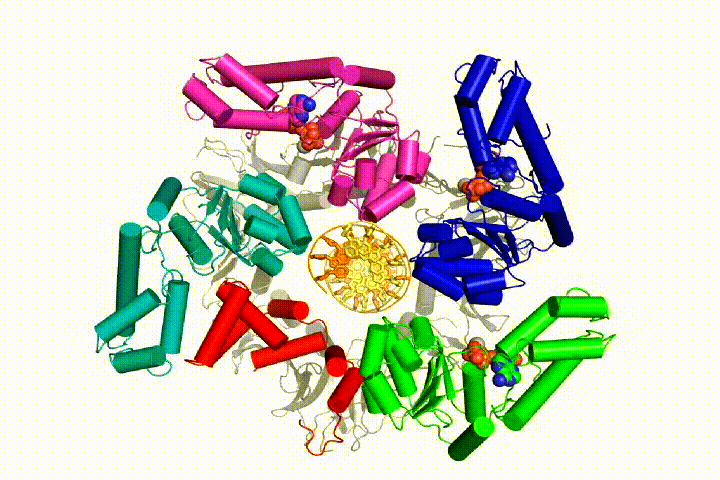
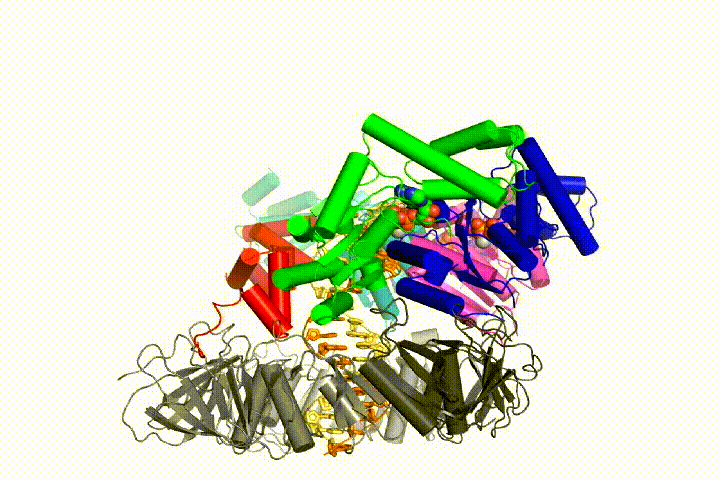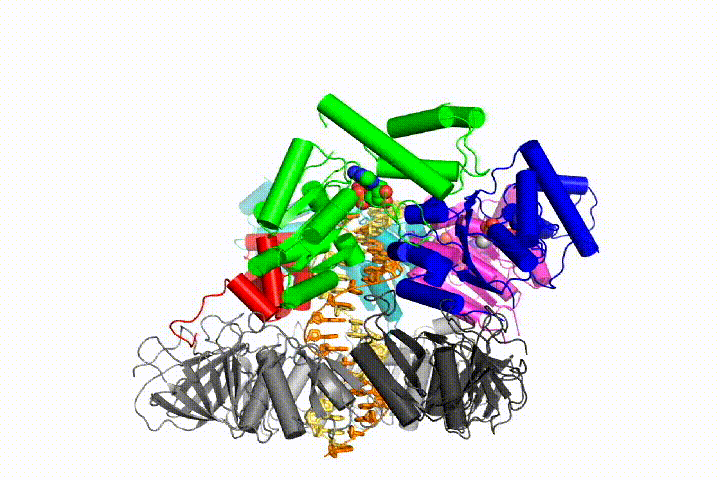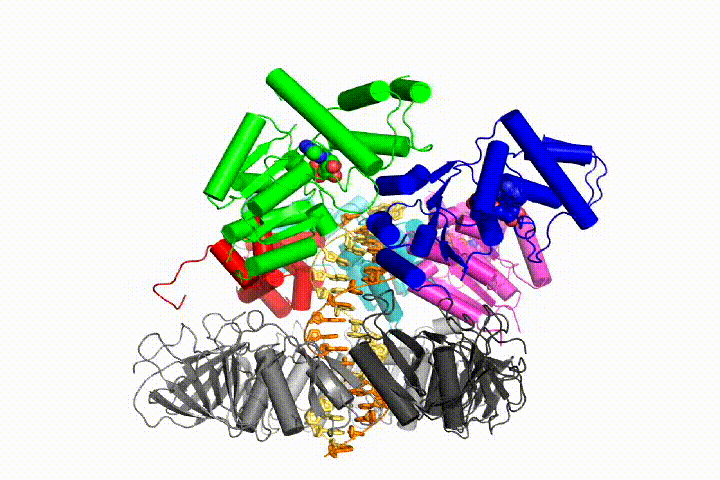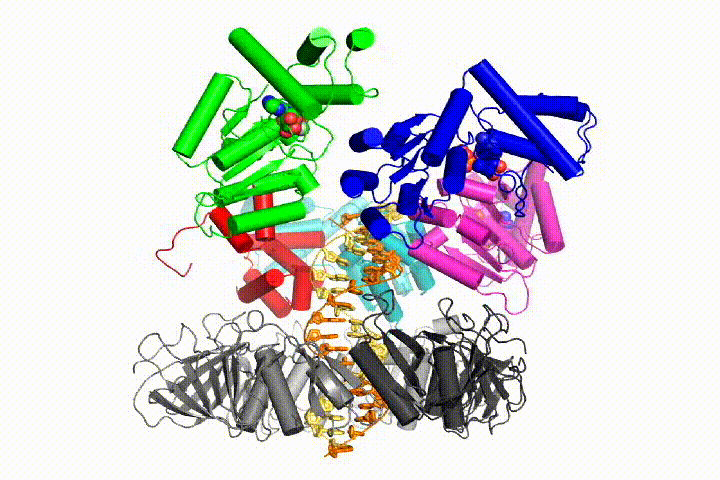
More recently, we have revealed the structural mechanism of clamp loading in unprecedented detail. Our cryo-EM structure of the human clamp loader RFC bound to PCNA revealed an early intermediate where the PCNA is closed and RFC is in an auto-inhibited conformation (Gaubitz & Liu et al PNAS 2020). This conformation is similar to a crystal structure of the yeast clamp loader, suggesting that this auto-inhibited state is conserved across a eukaryotes.
We more recently reported a detailed mechanism for the yeast clamp loader, with seven structures of clamp loading intermediates (Gaubitz et al, eLife 2022). We find a crab-claw mechanism for opening the clamp, which explains how the clamp loader achieves preference for its substrates. This allows DNA to directly bind into the large cleft, which induces clamp closure. We observe tight interactions at the ATPase active sites upon clamp closure, explaining how the clamp and DNA synergistically activate ATP hydrolysis and clamp release. More recently, we reported the discovery of a second DNA binding site on the exterior surface of the eukaryotic clamp loader (Liu & Gaubtiz et al eLife 2022). We reveal that both the external and internal DNA binding sites have the ability to partially unwind DNA without hydrolyzing ATP. Furthermore, we find that the two DNA binding sites cooperate to enable the clamp loader to bind to DNA at a nick or a small single-stranded gap, which are common intermediates during DNA repair. Finally, we show that yeast with a disrupted external DNA binding site have a defective DNA damage response, illustrating that this site is important for physiological function.
We test these models for clamp loader mechanism using a wide array of biochemical, biophysical and structural methods. Understanding the clamp loader mechanism in atomic detail will have important implications for understanding DNA replication as well as the mechanism of related AAA+ protein assemblies. Ultimately, we anticipate using our mechanistic understanding to develop novel nanodevices and small molecule effectors of AAA+ function. We have summarized our understanding of clamp loaders in reviews here, here and more recently here.

